Parts of this section have been published as:
Geissmann, T., 2002: Taxonomy and evolution of gibbons. In: Anthropology and primatology
into the third millennium: The Centenary Congress of the Zürich Anthropological
Institute (Evolutionary Anthropology Vol. 11, Supplement 1). Soligo, C.; Anzenberger,
G. & Martin, R.D. (eds.), pp. 28-31, New York: Wiley-Liss.
A printable pdf-file of this paper can be downloaded here.
Three different data sets of about equal size were collected in order to assess their relevance for a reconstruction of gibbon phylogeny with cladistic methods. Set 1 uses characteristics of fur coloration, set 2 consists of mainly morphological and anatomical data, and set 3 consists of vocal data.
Key words: gibbons; taxonomy; evolution; Hylobatidae; phylogeny; vocalizations; fur coloration.
INTRODUCTION |
Because the fossil history of gibbons is virtually
unknown (Fleagle, 1999; Geissmann, 1993), gibbon evolution can only be reconstructed
from a comparative analysis of evolutionarily informative characteristics of modern
gibbons and, to some degree, of related primate taxa that can be used as outgroups.
Several studies tried to reconstruct gibbon phylogeny, using fur coloration, morphological,
vocal or molecular data (Groves, 1972; Chivers, 1977; Haimoff et al., 1982; Creel
& Preuschoft, 1984; Garza & Woodruff, 1992; Hayashi et al., 1995; Purvis,
1995; Geissmann, 2002). Each study produced a different result. As a result, the
relationships among the various hylobatids are under debate and even the evolutionary
relationships among gibbon subgenera remain unresolved (Hall et al, 1998).
For the present study, three different data sets of about equal size were collected
in order to assess their relevance for a reconstruction of gibbon phylogeny with
cladistic methods. Set 1 uses characteristics of fur coloration, set 2 consists of
mainly morphological and anatomical data, and set 3 consists of vocal data.
MATERIALS AND METHODS |
Most recent studies agree that there are 4 distinct groups of gibbons, which are usually referred to as subgenera. Table 1 presents the classification used for the present study (Geissmann, 1995, 1997).
| Genus | Diploid number of chromosomes | Other division names | Species | |
| Hylobates |
44 |
lar group | H. agilis b | Agile gibbon |
| H. klossii | Kloss's gibbon | |||
| H. lar | White-handed gibbon | |||
| H. moloch | Silvery gibbon | |||
| H. muelleri c | Müller's gibbon | |||
| H. pileatus | Pileated gibbon | |||
| Bunopithecus |
38 |
B. hoolock | Hoolock | |
| Nomascus |
52 |
concolor group, crested gibbons | N. concolor | Western black crested gibbon |
| N. sp. cf. nasutus | Eastern black crested gibbon | |||
| N. gabriellae | Yellow-cheeked gibbon | |||
| N. leucogenys d | White-cheeked gibbon | |||
| Symphalangus |
50 |
S. syndactylus | Siamang |
a Raising the four main groups of gibbons to genus rank follows the consensus reached at the workshop "Primate Taxonomy for the New Millennium" (February 25-29, 2000, Orlando, Florida), to be presented in a future publication, and Roos & Geissmann, 2001.
b Including H. agilis albibarbis.
c Including H. muelleri abbotti and H. muelleri funereus.
d Including N. leucogenys siki.
I collected and analyzed three large data sets of
about equal size, as listed in Table 2. Data set were analyzed separately and compared
among each other with partition-homogeneity tests (Swofford, 2000).
Phylogenetic analysis were conducted with unweighted characters using the PAUP program
version 4.0b3a (Swofford, 2000). Two types of maximum parsimony trees were generated:
A branch-and-bound search of the same data set in PAUP yielded the shortest tree,
i.e. a single most parsimonious cladogram, and the bootstrap option of PAUP was used
to examine the robustness of internal nodes.
A hypothetical "ancestor" was used as an outgroup. This "ancestor"
was assembled using primitive character states wherever they could be reconstructed
or plausibly assumed.
The following standard measures were calculated in order to assess the "quality"
of trees: consistency index CI, retention index RI, and the rescaled consistency
index RC (Kitching et al, 1998; Maddison & Maddison, 1992), as well as the number
of bootstrap values larger than 50. The indices basically range from 0 to 1, with
higher RC values indicating that characters in the data set are more congruent with
each other and the tree.
Mean pairwise distances between taxa (i.e. the sum of their character state differences,
divided by the number of characters) were calculated with each data set (Swofford,
2000). Three types of distances were compared (1) distances among subspecies of the
same species (n=4), (2) distances among species of the same genus (n=38), and (3)
distances among members belonging to different genera (n=63). In order to increase
the sample of subspecies distances, H. muelleri was separated into its subspecies
(H. m. muelleri, H. m. funereus and H. m. abbotti) for this
part of the analysis.
RESULTS |
Because the three data sets are statistically different
(p=0.01) when compared with the partition-homogenetity test, they are not combined
in this study.
The trees based on fur coloration (Fig. 1a) strongly contradict all previously published
gibbon phylogenies (Groves, 1972; Chivers, 1977; Haimoff et al., 1982; Creel &
Preuschoft, 1984; Garza & Woodruff, 1992; Hayashi et al., 1995; Purvis, 1995;
Geissmann, in prep.). For instance, members of the lar group appear in the
most basal positions of the two shortest trees (not shown). The lar group
is not identified as a monophyletic group: Hylobates klossii is shown as the
sister taxon of the siamang (S. syndactylus), obviously because both gibbons
are completely black. In contrast, most previously published phylogenies identify
H. klossii as a member of the genus Hylobates, whereas the siamang
is placed in the genus Symphalangus.
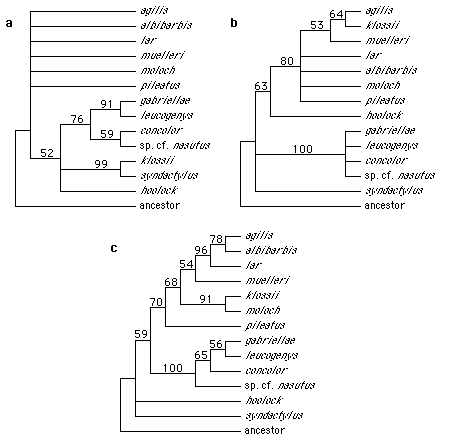
Figure 1. Maximum parsimony trees (bootstrap 50% majority-rule consensus) of three different data sets. a: Fur coloration data. b: "Non-communicatory" data. c: vocal data.
The trees based on "non-communicatory" data (Fig. 1b) resemble the trees
of earlier studies much more. The data set has a relatively low resolution because
of the many missing values in the data matrix. Many of the anatomical data were collected
during very early studies, when several gibbon taxa were simply not known. As a result
of the empty cells in the matrix, there is not one single most parsimonious tree,
but an ensemble of 117 shortest trees.
The trees based on vocal characteristics (Fig. 1c) are similar to those of data set
2. Bunopithecus hoolock appears in an unusually basal position in the tree,
however, because of its many apparently primitive vocal characteristics. Hylobates
klossii is identified as the sister taxon of H. moloch.
The various standard measures calculated in order to assess the "quality"
of trees are shown in Table 2 (bootstrap values, CI, RI, RC). Data set 1 scores distinctly
worse in all variables than the other two data sets. The difference between data
set 2 and 3 is less distinct in these variables. Nevertheless, data set 3 has higher
values in all of these variables (especially in the bootstrap values) than set 2,
indicating that particularly "good" trees are generated with vocal data.
| Data set |
Set 1 |
Set 2 |
Set 3 |
|||
| Number of characters |
37 |
34 |
34 |
|||
| Type of tree | S | B | S | B | S | B |
| Tree length | 158 | 191 | 73 | 78 | 88 | 90 |
| Number of bootstrap values above 50 | - | 5 | - | 5 | - | 1ß |
| Number of shortest trees | 2 | - | 117 | - | 1 | - |
| Consistency Index CI | 0.49 | 0.41 | 0.66 | 0.62 | 0.67 | 0.66 |
| Retention Index RI | 0.63 | 0.47 | 0.80 | 0.77 | 0.81 | 0.80 |
| Rescaled Consistency Index RC | 0.31 | 0.19 | 0.53 | 0.47 | 0.54 | 0.52 |
| Monophyly of agilis and albibarbis | - | - | - | - | + | + |
| Monophyly of concolor group | + | + | + | + | + | + |
| Monophyly of lar group (44-chromosome gibbons) | - | - | + | + | + | + |
| B. hoolock as sister group of lar group | - | - | + | + | - | - |
| H. klossii sister of all other members of lar group | - | - | - | - | - | - |
a Set 1 = Fur coloration data; Set 2 = "non-communicatory" data; Set 3 = vocal data. Abbreviations: S = shortest tree; B = Bootstrap 50% majority-rule consensus tree; + supported; - not supported
The mean pairwise distances between gibbon taxa are shown in Figure 2. Fur coloration
data are able to differentiate not only between species and genera, but also between
subspecies. Both the range and the standard deviation of the distances are extremely
large, however, and show a broad overlap among the different systematic levels. This
means that the differences in fur coloration provide little information on the genetic
distance between any two taxa. Two subspecies may be more different in fur coloration
than members of two different genera.
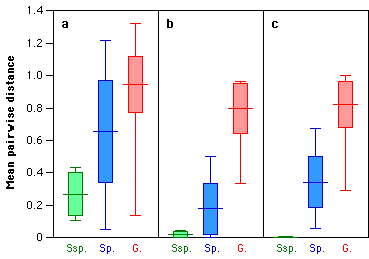
Figure 2. Pairwise character differences between (from left to right) different subspecies (Ssp.) of the same species, species (Sp.) of the same genus, and genera (G.) of the same family. a. Fur coloration data; b. "non-communicatory" data; c. vocal data. Each box plot shows the mean value (horizontal line through box), standard deviation (box) and range ("whiskers").
"Non-communicatory" data, on the other hand, do not differentiate well
between subspecies, and the standard deviations for subspecies and species overlap.
These data differentiate between some, but not all, species. The pairwise differences
between members of different genera are distinctly higher than those between species,
and standard deviations for species and genera do not overlap.
The vocal data produce similar results to the "non-communicatory" data.
This data set does not differentiate between subspecies at all, but differentiates
well between species and genera. Standard deviations of neither subspecies, species
or genera overlap.
The three data sets also differ in the species groups they support (Table 2). The
sets do agree, however, in supporting monophyly of the concolor group at least
to some degree, and in not supporting H. klossii
as the sister taxon to the other members of the lar group.
DISCUSSION |
A cladistic analysis suggests that the tempo of
evolutionary change differs among the data sets under study, similar to DNA sequences
derived from different parts of the genome. Fur coloration characters appear to change
considerably faster than each "non-communicatory" and vocal characters.
The three data sets produce different results and each set appears to be suited to
the analysis of different levels of resolution within the hylobatid radiation. Fur
coloration characteristics appear to provide little information for a gibbon phylogeny,
but may be valuable tools for subspecies identification, in contrast to most characters
of the other two data sets. In addition, many fur coloration characters differ among
individuals of the same taxon (polymorphisms).
"Non-communicatory" data appear to be much better suited for reconstructing
the gibbon phylogeny, but it is vocal data which produce the most reliable phylogeny
of the data sets under study. The trees generated with vocal data suggest:
- The gibbons of the genus Hylobates (lar group) and the genus Nomascus
(crested gibbons or concolor group) each are monophyletic groups. There is
weak support for a sister group relationship between the concolor and lar
groups.
- Hylobates klossii is neither the sister taxon of the siamang (S. syndactylus)
as suggested by some early studies (Elliot, 1913; Miller, 1903), nor the sister taxon
or the most basal group of the lar group, in contrast to many previous studies
(Chivers, 1977; Haimoff et al., 1982; Creel & Preuschoft, 1984; Purvis, 1995;
Schultz, 1933). This species is a fully integrated member of the lar group
(Garza & Woodruff, 1992) and apparently the sister taxon of H. moloch.
The same conclusion was independently reached by a study using a much smaller set
of vocal characteristics (including the degree of sex-specificity of the vocal repertoire,
the occurrence of solo songs and the preference for a specific time of day for song
production) (Geissmann, in prep.).
- Bunopithecus hoolock may be more basal than previously believed. Most earlier
studies recognized this species as the sister taxon to the lar group (Chivers,
1977; Haimoff et al., 1982; Purvis, 1995), whereas a more basal position has only
rarely been suggested (Creel & Preuschoft, 1984; Geissmann, in prep.).
The radiation of the four subgenera and of the species within the subgenera is not
reliably resolved with these preliminary results. In future studies, the reliability
of the trees may be improved by integrating data from several data sets, by weighting
the characters and by the supplementing the missing variables
in data set 2.
REFERENCES |
Geissmann,
T., 2002: Duet-splitting and the evolution of gibbon songs. Biological Reviews 77:
57-76.
Groves, CP. 1972. Systematics and phylogeny of gibbons. In: Rumbaugh, DM, editor.
Gibbon and siamang, vol. 1. Basel and New York: Karger. p 1-89.
Haimoff, EH, Chivers, DJ, Gittins, SP & Whitten, AJ. 1982. A phylogeny of gibbons
(Hylobates spp.) based on morphological and behavioural characters. Folia Primatol
39:213-237.
Hall, LM, Jones, DS & Wood, BA. 1998. Evolution of the gibbon subgenera inferred
from cytochrome b DNA sequence data. Molec Phylogenet Evol 10:281-286.
Hayashi, S, Hayasaka, K, Takenaka, O & Horai, S. 1995. Molecular phylogeny of
gibbons inferred from mitochondrial DNA sequences: Preliminary report. J Molec Evol
41:359-365.
Kitching, IJ, Forey, PL, Humphries, CJ & Williams, DM 1998. Cladistics. Second
edition. The theory and practice of parsimony analysis. Oxford and New York: Oxford
University Press.
Maddison, WP & Maddison, DR 1992. MacClade: Analysis of phylogeny and character
evolution. Version 3.0. Sunderland, Massachusetts: Sinauer Associates.
Miller, GS, Jr. 1903. Seventy new Malayan mammals. Smithson Misc Coll 45:1-73, +
19 plates.
Purvis, A. 1995. A composite estimate of primate phylogeny. Philosoph Transact Roy
Soc London B 348:405-421.
Roos,
C. & Geissmann, T., 2001. Molecular phylogeny of the major hylobatid divisions.
Molecular Phylogenetics and Evolution 19: 486-494.
Schultz, AH. 1933. Observations on the growth, classification and evolutionary specialization
of gibbons and siamangs. Hum Biol 5:212-255, and 385-428.
Swofford, DL. 2000. PAUP*. Phylogenetic analysis using parsimony (*and other methods).
Version 4. Sunderland, Massachusetts: Sinauer Associates
![]() .
.